An R6 class to represent an SBM fit with missing data
Source:R/R6Class-missSBM_fit.R
missSBM_fit.RdThe function estimateMissSBM() fits a collection of SBM for varying number of block.
Each fitted SBM is an instance of an R6 object with class missSBM_fit, described here.
Fields are accessed via active binding and cannot be changed by the user.
This class comes with a set of R6 methods, some of them being useful for the user and exported
as S3 methods. See the documentation for show(), print(), fitted(), predict(), plot().
Active bindings
fittedSBMthe fitted SBM with class
SimpleSBM_fit_noCov,SimpleSBM_fit_withCovorSimpleSBM_fit_MNARinheriting from classsbm::SimpleSBM_fitfittedSamplingthe fitted sampling, inheriting from class
networkSamplingand corresponding fitsimputedNetworkThe network data as a matrix with NAs values imputed with the current model
monitoringa list carrying information about the optimization process
entropyImputedthe entropy of the distribution of the imputed dyads
entropythe entropy due to the distribution of the imputed dyads and of the clustering
vExpecdouble: variational expectation of the complete log-likelihood
penaltydouble, value of the penalty term in ICL
loglikdouble: approximation of the log-likelihood (variational lower bound) reached
ICLdouble: value of the integrated classification log-likelihood
Methods
Method new()
constructor for networkSampling
Usage
missSBM_fit$new(partlyObservedNet, netSampling, clusterInit, useCov = TRUE)Arguments
partlyObservedNetAn object with class
partlyObservedNetwork.netSamplingThe sampling design for the modelling of missing data: MAR designs ("dyad", "node") and MNAR designs ("double-standard", "block-dyad", "block-node" ,"degree")
clusterInitInitial clustering: a vector with size
ncol(adjacencyMatrix), providing a user-defined clustering. The number of blocks is deduced from the number of levels in withclusterInit.useCovlogical. If covariates are present in partlyObservedNet, should they be used for the inference or of the network sampling design, or just for the SBM inference? default is TRUE.
Method doVEM()
a method to perform inference of the current missSBM fit with variational EM
Usage
missSBM_fit$doVEM(
control = list(threshold = 0.01, maxIter = 100, fixPointIter = 3, trace = TRUE)
)Arguments
controla list of parameters controlling the variational EM algorithm. See details of function
estimateMissSBM()
Examples
## Sample 75% of dyads in French political Blogosphere's network data
adjMatrix <- missSBM::frenchblog2007 %>%
igraph::as_adj (sparse = FALSE) %>%
missSBM::observeNetwork(sampling = "dyad", parameters = 0.75)
collection <- estimateMissSBM(adjMatrix, 3:5, sampling = "dyad")
#>
#>
#> Adjusting Variational EM for Stochastic Block Model
#>
#> Imputation assumes a 'dyad' network-sampling process
#>
#> Initialization of 3 model(s).
#> Performing VEM inference
#> Model with 5 blocks.
Model with 3 blocks.
Model with 4 blocks.
#> Looking for better solutions
#> Pass 1 Going forward ++
Pass 1 Going backward ++
my_missSBM_fit <- collection$bestModel
class(my_missSBM_fit)
#> [1] "missSBM_fit" "R6"
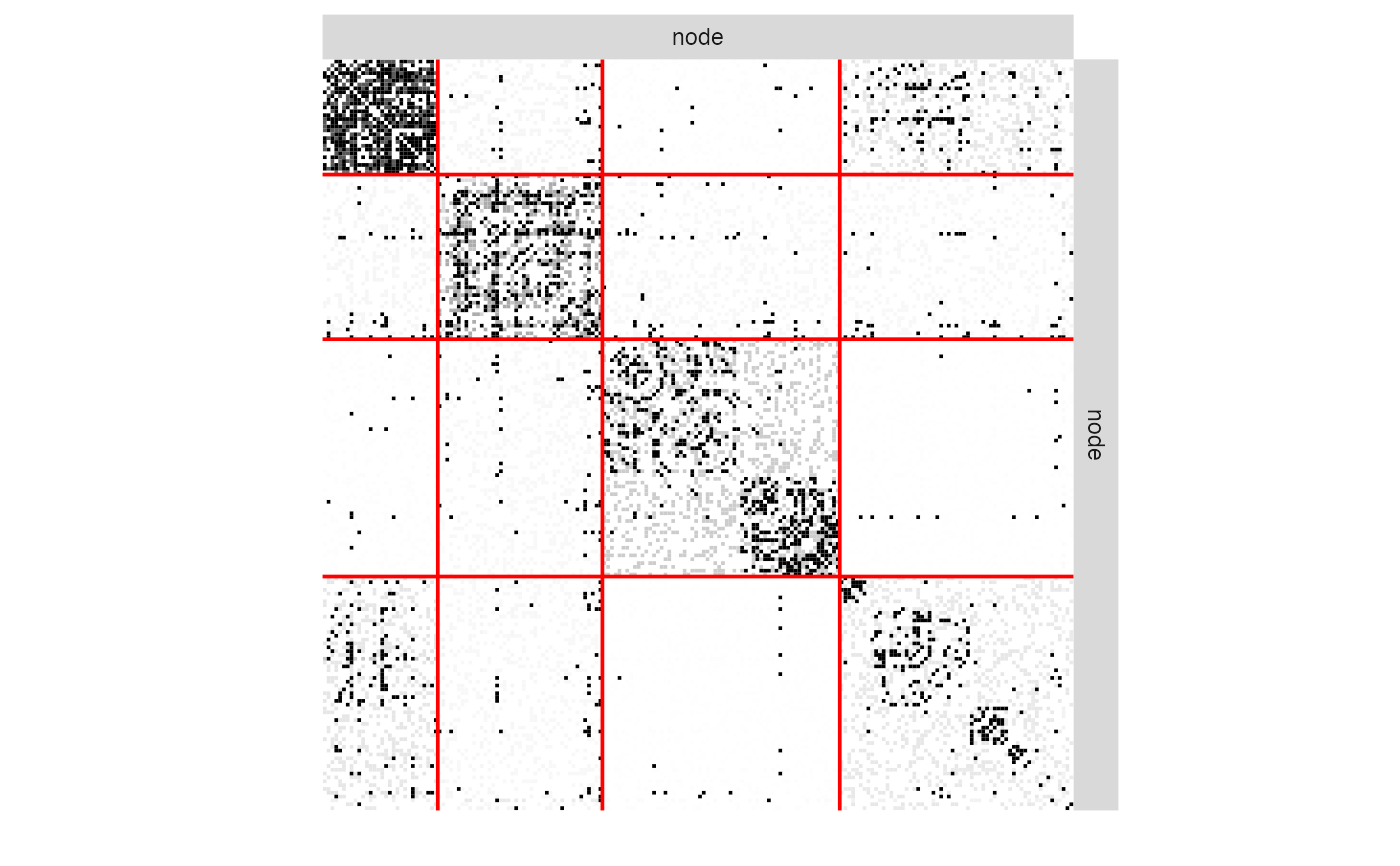
plot(my_missSBM_fit, "imputed")